
In a previous post we have covered how we can use Latent Growth Modeling in R to look at change in time. In that post we have assumed a simple linear model. This assumption is often unrealistic. Here I am going to show how we can free this assumption and find the best way to treat change in time.
We can use exploratory analysis and previous research as a starting point for understanding how to model change in time. Aditioanlly, we can also compare models that treat change in time in different ways to find the best fitting one for our data.
If we look again at the logincome in the Understanding Society data we get this graph (see previous post for an explanation of the data and syntax):
ggplot(usl, aes(wave, logincome, group = pidp)) +
geom_line(alpha = 0.01) + # add individual line with transparency
stat_summary( # add average line
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "red"
) +
theme_bw() + # nice theme
labs(x = "Wave", y = "Logincome") # nice labels
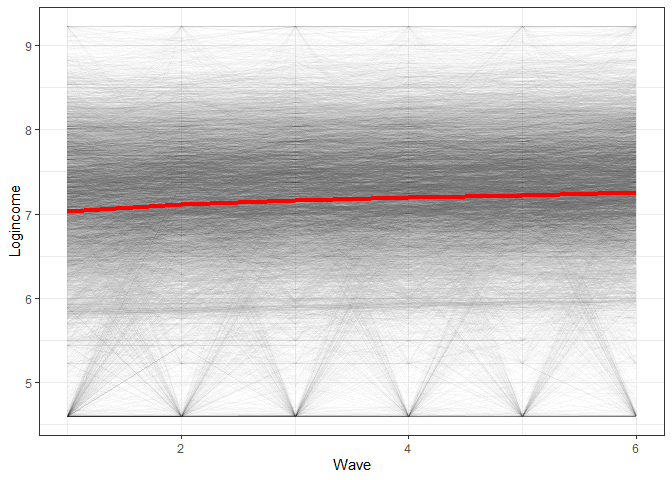
The graph would indicate we have an average change that is overall linear with a slight downward bend.
To see the individual level chenge I also sample 20 individuals and plot for each one their change in logincome.
# sample 20 ids people <- unique(usl$pidp) %>% sample(20) # do separate graph for each individual usl %>% filter(pidp %in% people) %>% # filter only sampled cases ggplot(aes(wave, logincome, group = 1)) + geom_line() + facet_wrap(~pidp) + # a graph for each individual theme_bw() + # nice theme labs(x = "Wave", y = "Logincome") # nice labels
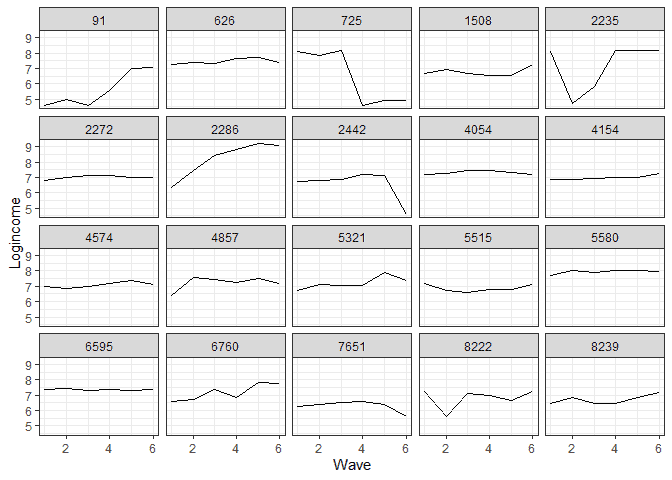
It appears that at the individual level we have a more mixed bag, although for quite a few people linear change would not be a bad approximation.
Keeping this in mind we can also decide on the best way to model change in time by comparing a number of different models and seeing which one fits the data best. As a starting point we can run the linear model which will be our reference (see previous post for an explanation of the model and syntax):
library(tidyverse)
library(lavaan)
# first LGM
model <- 'i =~ 1*logincome_1 + 1*logincome_2 + 1*logincome_3 +
1*logincome_4 + 1*logincome_5 + 1*logincome_6
s =~ 0*logincome_1 + 1*logincome_2 + 2*logincome_3 +
3*logincome_4 + 4*logincome_5 + 5*logincome_6'
fit1 <- growth(model, data = usw)
summary(fit1, standardized = TRUE)
## lavaan 0.6-8 ended normally after 45 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 11
##
## Number of observations 8752
##
## Model Test User Model:
##
## Test statistic 523.408
## Degrees of freedom 16
## P-value (Chi-square) 0.000
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## i =~
## logincome_1 1.000 0.827 0.842
## logincome_2 1.000 0.827 0.923
## logincome_3 1.000 0.827 0.949
## logincome_4 1.000 0.827 0.977
## logincome_5 1.000 0.827 0.993
## logincome_6 1.000 0.827 0.949
## s =~
## logincome_1 0.000 0.000 0.000
## logincome_2 1.000 0.115 0.128
## logincome_3 2.000 0.229 0.263
## logincome_4 3.000 0.344 0.406
## logincome_5 4.000 0.459 0.551
## logincome_6 5.000 0.574 0.658
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## i ~~
## s -0.047 0.002 -25.789 0.000 -0.497 -0.497
##
## Intercepts:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .logincome_1 0.000 0.000 0.000
## .logincome_2 0.000 0.000 0.000
## .logincome_3 0.000 0.000 0.000
## .logincome_4 0.000 0.000 0.000
## .logincome_5 0.000 0.000 0.000
## .logincome_6 0.000 0.000 0.000
## i 7.063 0.010 734.511 0.000 8.538 8.538
## s 0.041 0.002 23.527 0.000 0.354 0.354
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .logincome_1 0.280 0.006 45.748 0.000 0.280 0.291
## .logincome_2 0.201 0.004 48.502 0.000 0.201 0.250
## .logincome_3 0.212 0.004 54.736 0.000 0.212 0.279
## .logincome_4 0.197 0.004 54.733 0.000 0.197 0.275
## .logincome_5 0.176 0.004 49.002 0.000 0.176 0.254
## .logincome_6 0.217 0.005 43.584 0.000 0.217 0.286
## i 0.684 0.012 55.548 0.000 1.000 1.000
## s 0.013 0.000 31.248 0.000 1.000 1.000
Based on this model, on average, logincome is around 7.063 (~ £1,168) at the start of the study and goes up by 0.041 (~£1) each wave (more on the interpretation see previous post).
We can also look at change in time based on our model using visualizations (again, check previous post for explanations).
# predict the two latent variables
pred_lgm <- predict(fit1)
# create long data for each individual
pred_lgm_long <- map(0:5, # loop over time
function(x) pred_lgm[, 1] + x * pred_lgm[, 2]) %>%
reduce(cbind) %>% # bring together the wave predictions
as.data.frame() %>% # make data frame
setNames(str_c("Wave ", 1:6)) %>% # give names to variables
mutate(id = row_number()) %>% # make unique id
gather(-id, key = wave, value = pred) # make long format
# make graph (takes a minute to plot)
pred_lgm_long %>%
ggplot(aes(wave, pred, group = id)) + # what variables to plot?
geom_line(alpha = 0.01) + # add a transparent line for each person
stat_summary( # add average line
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "red"
) +
theme_bw() + # makes graph look nicer
labs(y = "logincome", # labels
x = "Wave")
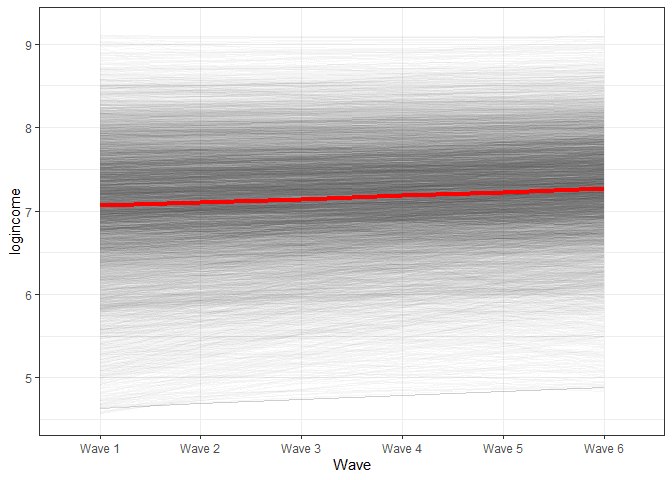
There are two general ways to expand this model to include non-linear change. One is by including polynomials while the other is by looking at relative change in time. We will cover both bellow.
Estimating non-linear LGM using polynomials
The inclusion of polynomials to model nonlinear effects has a similar motivation as in regression modeling. A polynomial (or interaction) allows the effect to change depending on the values of a predictor. In the case of LGM this would mean that we allow the slope to be higher or lower as time passes. This, in effect, would bend the trend upwards or downwards. If we want to allow for multiple bends then we need to include multiple polynomials. Bellow we will do this by including just the square effects modeled as a latent variable “q” (but the model can be easily expanded to include cubed effects and so on).
# square LGM
model <- 'i =~ 1*logincome_1 + 1*logincome_2 + 1*logincome_3 +
1*logincome_4 + 1*logincome_5 + 1*logincome_6
s =~ 0*logincome_1 + 1*logincome_2 + 2*logincome_3 +
3*logincome_4 + 4*logincome_5 + 5*logincome_6
q =~ 0*logincome_1 + 1*logincome_2 + 4*logincome_3 +
9*logincome_4 + 16*logincome_5 + 25*logincome_6'
fit2 <- growth(model, data = usw)
summary(fit2, standardized = TRUE)
## lavaan 0.6-8 ended normally after 93 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 15
##
## Number of observations 8752
##
## Model Test User Model:
##
## Test statistic 168.391
## Degrees of freedom 12
## P-value (Chi-square) 0.000
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## i =~
## logincome_1 1.000 0.843 0.877
## logincome_2 1.000 0.843 0.930
## logincome_3 1.000 0.843 0.956
## logincome_4 1.000 0.843 0.985
## logincome_5 1.000 0.843 1.002
## logincome_6 1.000 0.843 0.991
## s =~
## logincome_1 0.000 0.000 0.000
## logincome_2 1.000 0.271 0.299
## logincome_3 2.000 0.541 0.614
## logincome_4 3.000 0.812 0.949
## logincome_5 4.000 1.082 1.287
## logincome_6 5.000 1.353 1.590
## q =~
## logincome_1 0.000 0.000 0.000
## logincome_2 1.000 0.046 0.051
## logincome_3 4.000 0.185 0.210
## logincome_4 9.000 0.417 0.487
## logincome_5 16.000 0.741 0.881
## logincome_6 25.000 1.158 1.361
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## i ~~
## s -0.080 0.006 -12.637 0.000 -0.351 -0.351
## q 0.005 0.001 5.051 0.000 0.136 0.136
## s ~~
## q -0.011 0.001 -14.993 0.000 -0.893 -0.893
##
## Intercepts:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .logincome_1 0.000 0.000 0.000
## .logincome_2 0.000 0.000 0.000
## .logincome_3 0.000 0.000 0.000
## .logincome_4 0.000 0.000 0.000
## .logincome_5 0.000 0.000 0.000
## .logincome_6 0.000 0.000 0.000
## i 7.039 0.010 701.030 0.000 8.354 8.354
## s 0.071 0.005 14.233 0.000 0.261 0.261
## q -0.006 0.001 -6.317 0.000 -0.124 -0.124
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .logincome_1 0.213 0.008 26.812 0.000 0.213 0.231
## .logincome_2 0.207 0.004 49.387 0.000 0.207 0.252
## .logincome_3 0.196 0.004 50.313 0.000 0.196 0.253
## .logincome_4 0.178 0.004 48.924 0.000 0.178 0.243
## .logincome_5 0.178 0.004 49.022 0.000 0.178 0.252
## .logincome_6 0.173 0.007 25.732 0.000 0.173 0.239
## i 0.710 0.014 49.658 0.000 1.000 1.000
## s 0.073 0.004 16.951 0.000 1.000 1.000
## q 0.002 0.000 15.218 0.000 1.000 1.000
It appears that initially logincome goes up, by 0.071 each wave, but as time passes this bends downwards (because the effect is negative), by 0.006 each wave. The variance of “q” (0.002) is the between variation in non-linear change. Substantively, it tells us if people have different non-linear bends in time. If this would be 0 it means that everyone is following the same non-linear trend. If the value is large it means that individuals are very different in the non-linear part of the change in time of income.
Next, let’s plot the new estimates of change from the new model. We will use a similar procedure as above. The main change is to the formula. Now we need to add a new term which is time squared (x^2) multiplied by the coefficient for the square effect (pred_lgm2[, 3]). We also add in blue the line from the linear model for comparison.
# predict scores
pred_lgm2 <- predict(fit2)
# create long data for each individual
pred_lgm2_long <- map(0:5, # loop over time
function(x) pred_lgm2[, 1] +
x * pred_lgm2[, 2] +
x^2 * pred_lgm2[, 3]) %>%
reduce(cbind) %>% # bring together the wave predictions
as.data.frame() %>% # make data frame
setNames(str_c("Wave ", 1:6)) %>% # give names to variables
mutate(id = row_number()) %>% # make unique id
gather(-id, key = wave, value = pred) # make long format
# make graph
pred_lgm2_long %>%
ggplot(aes(wave, pred, group = id)) + # what variables to plot?
geom_line(alpha = 0.01) + # add a transparent line for each person
stat_summary( # add average line
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "blue"
) +
stat_summary(data = pred_lgm_long, # add average from linear model
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "red",
alpha = 0.5
) +
theme_bw() + # makes graph look nicer
labs(y = "logincome", # labels
x = "Wave")
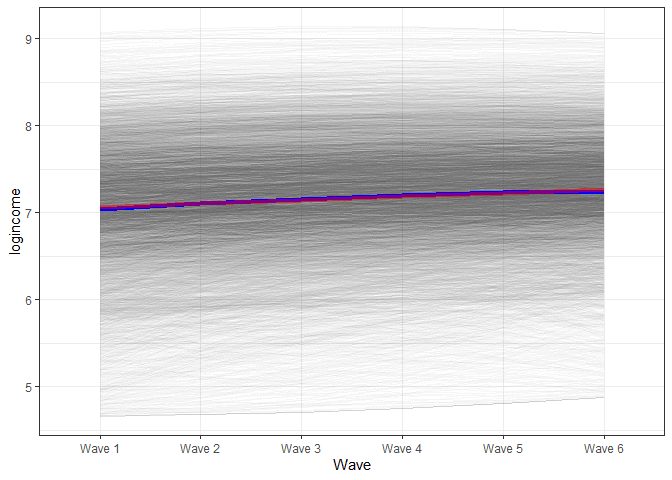
In the graph we see that the blue line (based on the model with the squared effects) starts slightly bellow the red line, by the middle is slightly above and in the end again, is slightly lower. That being said you really need to squint your eyes to see a difference. While the square effects are significant they might not be of substantive interest.
Non-linear change in time using relative change
The alternative way to model non-linear change is to estimate relative change. This is similar in spirit to including dummy variables in a regression model. The only thing we need to do is to tweak the loadings for the slope latent variable. Now we will fix only the first and the last loading to 0 and 1. The rest of the loadings will not be fixed and will be estimated. Now the interpretation of the slope will be the total amount of change from the first wave to the last one. The newly estimated loadings will tell us the proportion of change that happened from the start until that point out of the total change observed.
# relative change LGM
model <- 'i =~ 1*logincome_1 + 1*logincome_2 + 1*logincome_3 +
1*logincome_4 + 1*logincome_5 + 1*logincome_6
s =~ 0*logincome_1 + logincome_2 + logincome_3 +
logincome_4 + logincome_5 + 1*logincome_6'
fit3 <- growth(model, data = usw)
summary(fit3, standardized = TRUE)
## lavaan 0.6-8 ended normally after 113 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 15
##
## Number of observations 8752
##
## Model Test User Model:
##
## Test statistic 413.271
## Degrees of freedom 12
## P-value (Chi-square) 0.000
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## i =~
## logincome_1 1.000 0.821 0.838
## logincome_2 1.000 0.821 0.914
## logincome_3 1.000 0.821 0.939
## logincome_4 1.000 0.821 0.971
## logincome_5 1.000 0.821 0.988
## logincome_6 1.000 0.821 0.930
## s =~
## logincome_1 0.000 0.000 0.000
## logincome_2 0.158 0.023 6.977 0.000 0.081 0.091
## logincome_3 0.410 0.017 23.519 0.000 0.212 0.242
## logincome_4 0.718 0.017 43.392 0.000 0.370 0.438
## logincome_5 0.958 0.020 48.368 0.000 0.494 0.595
## logincome_6 1.000 0.516 0.585
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## i ~~
## s -0.202 0.011 -19.018 0.000 -0.477 -0.477
##
## Intercepts:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .logincome_1 0.000 0.000 0.000
## .logincome_2 0.000 0.000 0.000
## .logincome_3 0.000 0.000 0.000
## .logincome_4 0.000 0.000 0.000
## .logincome_5 0.000 0.000 0.000
## .logincome_6 0.000 0.000 0.000
## i 7.070 0.010 719.672 0.000 8.616 8.616
## s 0.174 0.008 20.993 0.000 0.337 0.337
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .logincome_1 0.286 0.008 37.707 0.000 0.286 0.298
## .logincome_2 0.189 0.005 37.392 0.000 0.189 0.235
## .logincome_3 0.211 0.004 53.437 0.000 0.211 0.277
## .logincome_4 0.193 0.004 53.163 0.000 0.193 0.270
## .logincome_5 0.159 0.004 37.667 0.000 0.159 0.231
## .logincome_6 0.243 0.005 44.710 0.000 0.243 0.312
## i 0.673 0.013 50.205 0.000 1.000 1.000
## s 0.266 0.013 21.250 0.000 1.000 1.000
So based on these results from wave 1 to wave 6 logincome has increased, on average, by 0.174. If we had truly linear change we would expect the loadings to increase at the same rate of 0.2 per wave (five steps going from wave 1 to six out of a total of change of 1 (the value of the last loading)). So the loading for “logincom_2” should be 0.2, the next, 0.4, and so on. We don’t really see that. We see that the most change happened between waves 3 and 4. Then, around 30% of the total change happened ((0.718 – 0.410) * 100). On the other hand, very little change happened between waves 5 and 6, less than 5%. This result would indicate that we don’t really have linear change. We can visualize the change using agian a graph.
To make the nice graph using the formula we need to extract the loadings. We can use the parameterestimates() command to do that:
parameterestimates(fit3) ## lhs op rhs est se z pvalue ci.lower ci.upper ## 1 i =~ logincome_1 1.000 0.000 NA NA 1.000 1.000 ## 2 i =~ logincome_2 1.000 0.000 NA NA 1.000 1.000 ## 3 i =~ logincome_3 1.000 0.000 NA NA 1.000 1.000 ## 4 i =~ logincome_4 1.000 0.000 NA NA 1.000 1.000 ## 5 i =~ logincome_5 1.000 0.000 NA NA 1.000 1.000 ## 6 i =~ logincome_6 1.000 0.000 NA NA 1.000 1.000 ## 7 s =~ logincome_1 0.000 0.000 NA NA 0.000 0.000 ## 8 s =~ logincome_2 0.158 0.023 6.977 0 0.113 0.202 ## 9 s =~ logincome_3 0.410 0.017 23.519 0 0.376 0.444 ## 10 s =~ logincome_4 0.718 0.017 43.392 0 0.685 0.750 ## 11 s =~ logincome_5 0.958 0.020 48.368 0 0.919 0.996 ## 12 s =~ logincome_6 1.000 0.000 NA NA 1.000 1.000 ## 13 logincome_1 ~~ logincome_1 0.286 0.008 37.707 0 0.271 0.300 ## 14 logincome_2 ~~ logincome_2 0.189 0.005 37.392 0 0.180 0.199 ## 15 logincome_3 ~~ logincome_3 0.211 0.004 53.437 0 0.203 0.219 ## 16 logincome_4 ~~ logincome_4 0.193 0.004 53.163 0 0.186 0.200 ## 17 logincome_5 ~~ logincome_5 0.159 0.004 37.667 0 0.151 0.168 ## 18 logincome_6 ~~ logincome_6 0.243 0.005 44.710 0 0.232 0.253 ## 19 i ~~ i 0.673 0.013 50.205 0 0.647 0.700 ## 20 s ~~ s 0.266 0.013 21.250 0 0.242 0.291 ## 21 i ~~ s -0.202 0.011 -19.018 0 -0.223 -0.181 ## 22 logincome_1 ~1 0.000 0.000 NA NA 0.000 0.000 ## 23 logincome_2 ~1 0.000 0.000 NA NA 0.000 0.000 ## 24 logincome_3 ~1 0.000 0.000 NA NA 0.000 0.000 ## 25 logincome_4 ~1 0.000 0.000 NA NA 0.000 0.000 ## 26 logincome_5 ~1 0.000 0.000 NA NA 0.000 0.000 ## 27 logincome_6 ~1 0.000 0.000 NA NA 0.000 0.000 ## 28 i ~1 7.070 0.010 719.672 0 7.051 7.089 ## 29 s ~1 0.174 0.008 20.993 0 0.158 0.190
With some manipulation we can extract just want we want:
# extract just th eloadings of the slopes loadings <- parameterestimates(fit3) %>% # get estimates filter(lhs == "s", op == "=~") %>% # filter the rows we want .[["est"]] # extract "est" variable # print result loadings ## [1] 0.0000000 0.1576471 0.4102628 0.7176219 0.9575655 1.0000000
We can follow a similar approach to the one before to create the long data with predicted scores from the LGM. The only difference is that we now loop over the loadings instead of the numbers 0 to 5:
# predict scores
pred_lgm3 <- predict(fit3)
# create long data for each individual
pred_lgm3_long <- map(loadings, # loop over time
function(x) pred_lgm3[, 1] +
x * pred_lgm3[, 2]) %>%
reduce(cbind) %>% # bring together the wave predictions
as.data.frame() %>% # make data frame
setNames(str_c("Wave ", 1:6)) %>% # give names to variables
mutate(id = row_number()) %>% # make unique id
gather(-id, key = wave, value = pred) # make long format
pred_lgm3_long %>%
ggplot(aes(wave, pred, group = id)) + # what variables to plot?
geom_line(alpha = 0.01) + # add a transparent line for each person
stat_summary( # add average line
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "green"
) +
stat_summary(data = pred_lgm_long, # add average from linear model
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "red",
alpha = 0.5
) +
stat_summary(data = pred_lgm2_long, # add average from squared model
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "blue",
alpha = 0.5
) +
theme_bw() + # makes graph look nicer
labs(y = "logincome", # labels
x = "Wave")
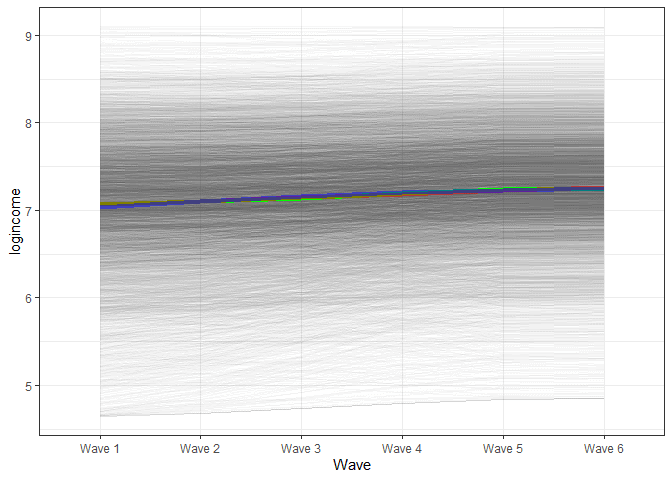
We see a very similar trend for the final model. It would appear that the trend here does not show a non-linear trajectory.
Finding the best fit
How to decide on the best model? We can compare relative fit indices, like AIC and BIC to help us with the decision (note that models are not nested so can’t use the Chi-squared test):
anova(fit1, fit2, fit3) ## Chi-Squared Difference Test ## ## Df AIC BIC Chisq Chisq diff Df diff Pr(>Chisq) ## fit2 12 97516 97622 168.39 ## fit3 12 97761 97867 413.27 244.88 0 ## fit1 16 97863 97941 523.41 110.14 4 < 2.2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Based on AIC and BIC the best fitting model is the one with square effects. That being said it’s always important to think if those effects are really important from a substantive point of view and if doing just a linear model would really lead to different conclusions.
Conclusions
Hopefully that gives you an idea how to estimate non-linear LGM, how to estimate it in R and how to visualize this change. It’s always useful to visualize these models as the interpretation can get quite tricky.
If you liked this you can have a look at training that I offer as well as other blog posts, such as this introduction to multilevel modeling for longitudinal data or this one looking at visualizing transition in time for categorical variables.

If that was useful you might also like the Longitudinal Data Analysis Using R book.
This covers everything you need to work with longitudinal data. It introduces the key concepts related to longitudinal data, the basics of R and regression. It also shows using real data how to prepare, explore and visualize longitudinal data. In addition, it discusses in depth popular statistical models such as the multilevel model for change, the latent growth model and the cross-lagged model.


Dear Alexandru,
Your code for latent growth curve modelling has been of great help in my work. I have a question concerning this post. You write that “we will do this by including just the square effects modeled as a latent variable “q” (but the model can be easily expanded to include cubed effects and so on)”
My question is: how can I write the code to include cubed effects, that is, to create a cubic model?
Thank you so much for sharing your work.
Best,
Gustaf
Thank you for the question. The code should be something like this. To run a model with cube you could use such a code (sorry the formatting is not great for some reason):
# cube Latent Growth Model
model <- 'i =~ 1*logincome_1 + 1*logincome_2 + 1*logincome_3 + 1*logincome_4 + 1*logincome_5 + 1*logincome_6 s =~ 0*logincome_1 + 1*logincome_2 + 2*logincome_3 + 3*logincome_4 + 4*logincome_5 + 5*logincome_6 q =~ 0*logincome_1 + 1*logincome_2 + 4*logincome_3 + 9*logincome_4 + 16*logincome_5 + 25*logincome_6 c =~ 0*logincome_1 + 1*logincome_2 + 8*logincome_3 + 27*logincome_4 + 64*logincome_5 + 125*logincome_6'
fit3 <- growth(model, data = usw)
To run the graph it should be something like this:
# predict scores
pred_lgm3 <- predict(fit3)
# create long data for each individual
pred_lgm3_long <- map(0:5, # loop over time function(x) pred_lgm2[, 1] + x * pred_lgm2[, 2] + x^2 * pred_lgm2[, 3] + x^3 * pred_lgm2[, 3]) %>%
reduce(cbind) %>% # bring together the wave predictions
as.data.frame() %>% # make data frame
setNames(str_c("Wave ", 1:6)) %>% # give names to variables
mutate(id = row_number()) %>% # make unique id
gather(-id, key = wave, value = pred) # make long format
# make graph
pred_lgm3_long %>%
ggplot(aes(wave, pred, group = id)) + # what variables to plot?
geom_line(alpha = 0.01) + # add a transparent line for each person
stat_summary( # add average line from cube model
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "blue"
) +
stat_summary(data = pred_lgm_long, # add average from linear model (optional)
aes(group = 1),
fun = mean,
geom = "line",
size = 1.5,
color = "red",
alpha = 0.5
) +
theme_bw() + # makes graph look nicer
labs(y = "logincome", # labels
x = "Wave")
Pingback: Estimating non-linear change in time using the multilevel model for change - Alexandru Cernat